|
 |
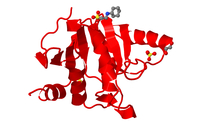
 3D structures of proteins and compounds related to Coronavirus SARS-CoV-2 |
 |
 |
 |
 |
 3D structures of proteins and compounds related to Coronavirus SARS-CoV-2 |
 |
 |
 "Searching Contact Molecules with Query Protein" of HOMCOS for each protein of the virus.
Monomeric and bound
"Searching Contact Molecules with Query Protein" of HOMCOS for each protein of the virus.
Monomeric and bound  3D structures of the virus proteins and their homologs are summarized. Found 3D structures can be used for templates of homology modeling. Details of the usage are explained in the HELP page.
3D structures of the virus proteins and their homologs are summarized. Found 3D structures can be used for templates of homology modeling. Details of the usage are explained in the HELP page.
| AC | ID | length (amino acids) |
Number of 3D homologues |
short name | full name |
 P0DTC3 P0DTC3 |
AP3A_SARS2 | 275 | 12 | ORF3a; | ORF3a protein; Accessory protein 3a;Protein 3a;Protein U274;Protein X1; |
 P0DTC9 P0DTC9 |
NCAP_SARS2 | 419 | 365 | N; NC ;Protein N ; | Nucleoprotein ; Nucleocapsid protein ; |
 P0DTC6 P0DTC6 |
NS6_SARS2 | 61 | 0 | ORF6; ns6; | ORF6 protein; Accessory protein 6;Non-structural protein 6;Protein X3; |
 P0DTC7 P0DTC7 |
NS7A_SARS2 | 121 | 4 | ORF7a; | ORF7a protein; Accessory protein 7a;Protein U122;Protein X4; |
 P0DTD8 P0DTD8 |
NS7B_SARS2 | 43 | 0 | ORF7b; | ORF7b protein; Accessory protein 7b; |
 P0DTC8 P0DTC8 |
NS8_SARS2 | 121 | 9 | ORF8; ns8; | ORF8 protein; Non-structural protein 8; |
 P0DTF1 P0DTF1 |
ORF3B_SARS2 | 22 | 0 | ORF3b; | Putative ORF3b protein ; |
 P0DTG1 P0DTG1 |
ORF3C_SARS2 | 41 | 0 | ORF3c; ORF3h; | ORF3c protein ; ORF3h protein ; |
 P0DTG0 P0DTG0 |
ORF3D_SARS2 | 57 | 0 | Putative ORF3d protein ; | |
 P0DTD2 P0DTD2 |
ORF9B_SARS2 | 97 | 24 | ORF9b; | ORF9b protein; Accessory protein 9b;ORF-9b;Protein 9b; |
 P0DTD3 P0DTD3 |
ORF9C_SARS2 | 73 | 0 | ORF9c; ORF14; | Putative ORF9c protein ; Uncharacterized protein 14; |
 P0DTD1 P0DTD1 |
R1AB_SARS2 | 7096 | 1500 | pp1ab; | Replicase polyprotein 1ab; ORF1ab polyprotein; |
 P0DTC1 P0DTC1 |
R1A_SARS2 | 4405 | 1500 | pp1a; | Replicase polyprotein 1a; ORF1a polyprotein; |
 P0DTC2 P0DTC2 |
SPIKE_SARS2 | 1273 | 1500 | S glycoprotein ; | Spike glycoprotein ; E2 ;Peplomer protein ; |
 P0DTC4 P0DTC4 |
VEMP_SARS2 | 75 | 18 | E;sM protein ; | Envelope small membrane protein ; |
 P0DTC5 P0DTC5 |
VME1_SARS2 | 222 | 20 | M; | Membrane protein ; E1 glycoprotein ;Matrix glycoprotein ;Membrane glycoprotein ; |
 P59632 P59632 |
AP3A_SARS | 274 | 12 | ORF3a protein; Accessory protein 3a;Protein 3a;Protein U274;Protein X1; | |
 P59595 P59595 |
NCAP_SARS | 422 | 366 | NC ;Protein N ; | Nucleoprotein ; Nucleocapsid protein ; |
 P59633 P59633 |
NS3B_SARS | 154 | 0 | ns3b; | ORF3b protein; Accessory protein 3b;Non-structural protein 3b;Protein X2; |
 P59634 P59634 |
NS6_SARS | 63 | 0 | ORF6; ns6; | ORF6 protein; Accessory protein 6;Non-structural protein 6;Protein X3; |
 P59635 P59635 |
NS7A_SARS | 122 | 4 | ORF7a protein; Accessory protein 7a;Protein U122;Protein X4; | |
 Q7TFA1 Q7TFA1 |
NS7B_SARS | 44 | 0 | ns7b; | Protein non-structural 7b; Accessory protein 7b; |
 Q7TFA0 Q7TFA0 |
NS8A_SARS | 39 | 0 | ns8a; | ORF8a protein; Accessory protein 8a;Protein non-structural 8a; |
 Q80H93 Q80H93 |
NS8B_SARS | 84 | 0 | ns8b; | ORF8b protein; Accessory protein 8b;Non-structural protein 8b; |
 P59636 P59636 |
ORF9B_SARS | 98 | 24 | ORF9b protein; Accessory protein 9b;ORF-9b;Protein 9b; | |
 P0C6X7 P0C6X7 |
R1AB_SARS | 7073 | 1500 | pp1ab; | Replicase polyprotein 1ab; ORF1ab polyprotein; |
 P0C6U8 P0C6U8 |
R1A_SARS | 4382 | 1500 | pp1a; | Replicase polyprotein 1a; ORF1a polyprotein; |
 P59594 P59594 |
SPIKE_SARS | 1255 | 1500 | S glycoprotein ; | Spike glycoprotein ; E2 ;Peplomer protein ; |
 P59637 P59637 |
VEMP_SARS | 76 | 18 | E protein ;sM protein ; | Envelope small membrane protein ; |
 P59596 P59596 |
VME1_SARS | 221 | 20 | M protein ; | Membrane protein ; E1 glycoprotein ;Matrix glycoprotein ;Membrane glycoprotein ; |
 Q7TLC7 Q7TLC7 |
Y14_SARS | 70 | 0 | Uncharacterized protein 14; | |
 Q9BYF1 Q9BYF1 |
ACE2_HUMAN | 805 | 796 | ACEH ;ACE-related carboxypeptidase; | Angiotensin-converting enzyme 2; Angiotensin-converting enzyme homolog ;Angiotensin-converting enzyme-related carboxypeptidase ;Metalloprotease MPROT15 {ECO:0000303|Ref.6}; |
 P30556 P30556 |
AGTR1_HUMAN | 359 | 1450 | AT1 receptor ; | Type-1 angiotensin II receptor ; AT1AR ;AT1BR;Angiotensin II type-1 receptor ; |
 Q4KMQ2 Q4KMQ2 |
ANO6_HUMAN | 910 | 131 | SCAN channel; | Anoctamin-6 ; Small-conductance calcium-activated nonselective cation channel;Transmembrane protein 16F; |
 P02649 P02649 |
APOE_HUMAN | 317 | 40 | Apo-E; | Apolipoprotein E ; |
 Q9NVJ2 Q9NVJ2 |
ARL8B_HUMAN | 186 | 1500 | ADP-ribosylation factor-like protein 8B ; ADP-ribosylation factor-like protein 10C;Novel small G protein indispensable for equal chromosome segregation 1; | |
 P17405 P17405 |
ASM_HUMAN | 631 | 35 | aSMase; | Sphingomyelin phosphodiesterase ; Acid sphingomyelinase {ECO:0000303|PubMed:25339683, ECO:0000303|PubMed:27349982, ECO:0000303|PubMed:33163980}; |
 P35613 P35613 |
BASI_HUMAN | 385 | 68 | EMMPRIN;HAb18G ;TCSF; | Basigin ; 5F7;Collagenase stimulatory factor;Extracellular matrix metalloproteinase inducer;Hepatoma-associated antigen ;Leukocyte activation antigen M6;OK blood group antigen;Tumor cell-derived collagenase stimulatory factor; |
 O00327 O00327 |
BMAL1_HUMAN | 626 | 192 | bHLHe5; | Basic helix-loop-helix ARNT-like protein 1 ; Aryl hydrocarbon receptor nuclear translocator-like protein 1;Basic-helix-loop-helix-PAS protein MOP3;Brain and muscle ARNT-like 1;Class E basic helix-loop-helix protein 5;Member of PAS protein 3;PAS domain-containing protein 3;bHLH-PAS protein JAP3; |
 Q10589 Q10589 |
BST2_HUMAN | 180 | 47 | BST-2; | Bone marrow stromal antigen 2; HM1.24 antigen;Tetherin; |
 P33076 P33076 |
C2TA_HUMAN | 1130 | 106 | CIITA ; | MHC class II transactivator ; |
 P07711 P07711 |
CATL1_HUMAN | 333 | 894 | MEP; | Procathepsin L ; Cathepsin L1 ;Major excreted protein; |
 O95992 O95992 |
CH25H_HUMAN | 272 | 0 | h25OH; | Cholesterol 25-hydroxylase ; Cholesterol 25-monooxygenase; |
 Q92499 Q92499 |
DDX1_HUMAN | 740 | 870 | DBP-RB; | ATP-dependent RNA helicase DDX1; DEAD box protein 1;DEAD box protein retinoblastoma; |
 O95786 O95786 |
RIGI_HUMAN | 925 | 230 | RLR-1;RIG-1;RIG-I; | Antiviral innate immune response receptor RIG-I ; ATP-dependent RNA helicase DDX58 ;DEAD box protein 58;RIG-I-like receptor 1;RNA sensor RIG-I ;Retinoic acid-inducible gene 1 protein;Retinoic acid-inducible gene I protein; |
 Q96C10 Q96C10 |
DHX58_HUMAN | 678 | 237 | RLR-3;RLR; | ATP-dependent RNA helicase DHX58 ; ATP-dependent helicase LGP2 ;Protein D11Lgp2 homolog;RIG-I-like receptor 3;RIG-I-like receptor LGP2; |
 P68104 P68104 |
EF1A1_HUMAN | 462 | 503 | EF-1-alpha-1; EF-Tu;eEF1A-1; | Elongation factor 1-alpha 1; Elongation factor Tu;Eukaryotic elongation factor 1 A-1;Leukocyte receptor cluster member 7; |
 P49327 P49327 |
FAS_HUMAN | 2511 | 1156 | Fatty acid synthase; Type I fatty acid synthase; | |
 P09958 P09958 |
FURIN_HUMAN | 794 | 306 | PACE; | Furin ; Dibasic-processing enzyme;Paired basic amino acid residue-cleaving enzyme; |
 Q9Y2I7 Q9Y2I7 |
FYV1_HUMAN | 2098 | 568 | Phosphatidylinositol 3-phosphate 5-kinase; PIPkin-III;Type III PIP kinase; | 1-phosphatidylinositol 3-phosphate 5-kinase ; FYVE finger-containing phosphoinositide kinase;PIKfyve;Phosphatidylinositol 3-phosphate 5-kinase type III;Serine-protein kinase PIKFYVE; |
 P04233 P04233 |
HG2A_HUMAN | 296 | 46 | Ii; | HLA class II histocompatibility antigen gamma chain ; HLA-DR antigens-associated invariant chain;Ia antigen-associated invariant chain; |
 Q16665 Q16665 |
HIF1A_HUMAN | 826 | 191 | HIF-1-alpha ;HIF1-alpha ; bHLHe78; | Hypoxia-inducible factor 1-alpha ; ARNT-interacting protein;Basic-helix-loop-helix-PAS protein MOP1 ;Class E basic helix-loop-helix protein 78;Member of PAS protein 1 ;PAS domain-containing protein 8; |
 P04439 P04439 |
HLAA_HUMAN | 365 | 1500 | HLA-A; | HLA class I histocompatibility antigen, A alpha chain; Human leukocyte antigen A; |
 P01889 P01889 |
HLAB_HUMAN | 362 | 1500 | HLA-B; | HLA class I histocompatibility antigen, B alpha chain; Human leukocyte antigen B; |
 P13747 P13747 |
HLAE_HUMAN | 358 | 1500 | HLA class I histocompatibility antigen, alpha chain E; MHC class I antigen E; | |
 P09429 P09429 |
HMGB1_HUMAN | 215 | 81 | HMG-1; | High mobility group protein B1; High mobility group protein 1; |
 Q96F46 Q96F46 |
I17RA_HUMAN | 866 | 28 | IL-17 receptor A;IL-17RA; | Interleukin-17 receptor A ; CDw217; |
 Q8NAC3 Q8NAC3 |
I17RC_HUMAN | 791 | 6 | IL-17 receptor C;IL-17RC; IL17Rhom;IL-17RL; | Interleukin-17 receptor C; Interleukin-17 receptor homolog;Interleukin-17 receptor-like protein;ZcytoR14; |
 Q9BYX4 Q9BYX4 |
IFIH1_HUMAN | 1025 | 332 | CADM-140 autoantigen;Helicard;MDA-5;RLR-2; | Interferon-induced helicase C domain-containing protein 1 ; Clinically amyopathic dermatomyositis autoantigen 140 kDa;Helicase with 2 CARD domains;Interferon-induced with helicase C domain protein 1;Melanoma differentiation-associated protein 5;Murabutide down-regulated protein;RIG-I-like receptor 2;RNA helicase-DEAD box protein 116; |
 P13164 P13164 |
IFM1_HUMAN | 125 | 0 | DSPA2a; | Interferon-induced transmembrane protein 1 ; Dispanin subfamily A member 2a;Interferon-induced protein 17;Interferon-inducible protein 9-27;Leu-13 antigen; |
 Q01629 Q01629 |
IFM2_HUMAN | 132 | 0 | DSPA2c; | Interferon-induced transmembrane protein 2 ; Dispanin subfamily A member 2c;Interferon-inducible protein 1-8D; |
 Q01628 Q01628 |
IFM3_HUMAN | 133 | 0 | DSPA2b; | Interferon-induced transmembrane protein 3 ; Dispanin subfamily A member 2b;Interferon-inducible protein 1-8U; |
 Q96PD4 Q96PD4 |
IL17F_HUMAN | 163 | 106 | IL-17F; | Interleukin-17F; Cytokine ML-1; |
 Q16552 Q16552 |
IL17_HUMAN | 155 | 100 | IL-17;IL-17A; CTLA-8; | Interleukin-17A; Cytotoxic T-lymphocyte-associated antigen 8; |
 P08887 P08887 |
IL6RA_HUMAN | 468 | 118 | IL-6 receptor subunit alpha;IL-6R subunit alpha;IL-6R-alpha;IL-6RA; gp80; | Interleukin-6 receptor subunit alpha ; IL-6R 1;Membrane glycoprotein 80; |
 P40189 P40189 |
IL6RB_HUMAN | 918 | 183 | IL-6 receptor subunit beta;IL-6R subunit beta;IL-6R-beta;IL-6RB; gp130 ; | Interleukin-6 receptor subunit beta ; CDw130;Interleukin-6 signal transducer;Membrane glycoprotein 130;Oncostatin-M receptor subunit alpha; |
 P05231 P05231 |
IL6_HUMAN | 212 | 26 | IL-6; BSF-2;CDF;IFN-beta-2; | Interleukin-6 ; B-cell stimulatory factor 2;CTL differentiation factor;Hybridoma growth factor;Interferon beta-2; |
 P52292 P52292 |
IMA1_HUMAN | 529 | 531 | Importin subunit alpha-1 ; Karyopherin subunit alpha-2;RAG cohort protein 1;SRP1-alpha; | |
 P17181 P17181 |
INAR1_HUMAN | 557 | 49 | IFN-R-1;IFN-alpha/beta receptor 1; CRF2-1; | Interferon alpha/beta receptor 1; Cytokine receptor class-II member 1;Cytokine receptor family 2 member 1;Type I interferon receptor 1; |
 P48551 P48551 |
INAR2_HUMAN | 515 | 12 | IFN-R-2;IFN-alpha binding protein;IFN-alpha/beta receptor 2; | Interferon alpha/beta receptor 2; Interferon alpha binding protein;Type I interferon receptor 2; |
 Q14653 Q14653 |
IRF3_HUMAN | 427 | 101 | IRF-3 ; | Interferon regulatory factor 3 {ECO:0000303|PubMed:9566918, ECO:0000303|PubMed:9803267}; |
 Q13568 Q13568 |
IRF5_HUMAN | 498 | 103 | IRF-5 ; | Interferon regulatory factor 5 ; |
 Q92985 Q92985 |
IRF7_HUMAN | 503 | 101 | IRF-7; | Interferon regulatory factor 7; |
 P05161 P05161 |
ISG15_HUMAN | 165 | 1500 | IP17;hUCRP; | Ubiquitin-like protein ISG15; Interferon-induced 15 kDa protein;Interferon-induced 17 kDa protein;Ubiquitin cross-reactive protein; |
 P20701 P20701 |
ITAL_HUMAN | 1170 | 433 | LFA-1A; | Integrin alpha-L ; CD11 antigen-like family member A;Leukocyte adhesion glycoprotein LFA-1 alpha chain;Leukocyte function-associated molecule 1 alpha chain; |
 Q13241 Q13241 |
KLRD1_HUMAN | 179 | 619 | Natural killer cells antigen CD94; KP43;Killer cell lectin-like receptor subfamily D member 1;NK cell receptor; | |
 Q16553 Q16553 |
LY6E_HUMAN | 131 | 0 | Ly-6E; RIG-E;SCA-2;TSA-1; | Lymphocyte antigen 6E ; Retinoic acid-induced gene E protein;Stem cell antigen 2;Thymic shared antigen 1; |
 Q7Z434 Q7Z434 |
MAVS_HUMAN | 540 | 72 | MAVS ; Cardif ;IPS-1;VISA; | Mitochondrial antiviral-signaling protein ; CARD adapter inducing interferon beta ;Interferon beta promoter stimulator protein 1;Putative NF-kappa-B-activating protein 031N;Virus-induced-signaling adapter; |
 Q86U44 Q86U44 |
MTA70_HUMAN | 580 | 160 | hMETTL3 ;MT-A70; | N(6)-adenosine-methyltransferase catalytic subunit METTL3 ; Methyltransferase-like protein 3 ;N(6)-adenosine-methyltransferase 70 kDa subunit; |
 Q99836 Q99836 |
MYD88_HUMAN | 296 | 277 | Myeloid differentiation primary response protein MyD88 ; | |
 Q9Y6K9 Q9Y6K9 |
NEMO_HUMAN | 419 | 61 | NEMO ; IKKAP1;I-kappa-B kinase subunit gamma;IKK-gamma;IKKG;IkB kinase subunit gamma; | NF-kappa-B essential modulator ; FIP-3;IkB kinase-associated protein 1;Inhibitor of nuclear factor kappa-B kinase subunit gamma;NF-kappa-B essential modifier; |
 P01185 P01185 |
NEU2_HUMAN | 164 | 29 | Vasopressin-neurophysin 2-copeptin; AVP-NPII; | |
 Q16236 Q16236 |
NF2L2_HUMAN | 605 | 12 | NF-E2-related factor 2 ;NFE2-related factor 2 ;Nrf-2 ; | Nuclear factor erythroid 2-related factor 2 ; Nuclear factor, erythroid derived 2, like 2 {ECO:0000303|PubMed:33009401, ECO:0000303|PubMed:7937919}; |
 O14745 O14745 |
NHRF1_HUMAN | 358 | 325 | NHERF-1; EBP50; | Na(+)/H(+) exchange regulatory cofactor NHE-RF1 ; Ezrin-radixin-moesin-binding phosphoprotein 50;Regulatory cofactor of Na(+)/H(+) exchanger;Sodium-hydrogen exchanger regulatory factor 1;Solute carrier family 9 isoform A3 regulatory factor 1; |
 P26715 P26715 |
NKG2A_HUMAN | 233 | 460 | NKG2-A/NKG2-B type II integral membrane protein; CD159 antigen-like family member A;NK cell receptor A;NKG2-A/B-activating NK receptor; | |
 Q9C000 Q9C000 |
NLRP1_HUMAN | 1473 | 294 | NACHT, LRR and PYD domains-containing protein 1 ; Caspase recruitment domain-containing protein 7;Death effector filament-forming ced-4-like apoptosis protein ;Nucleotide-binding domain and caspase recruitment domain; | |
 Q96P20 Q96P20 |
NLRP3_HUMAN | 1036 | 222 | CLR1.1 ; | NACHT, LRR and PYD domains-containing protein 3 ; Angiotensin/vasopressin receptor AII/AVP-like;Caterpiller protein 1.1 ;Cold-induced autoinflammatory syndrome 1 protein ;Cryopyrin ;PYRIN-containing APAF1-like protein 1 ; |
 O14786 O14786 |
NRP1_HUMAN | 923 | 246 | Neuropilin-1 ; Vascular endothelial cell growth factor 165 receptor; | |
 P52948 P52948 |
NUP98_HUMAN | 1817 | 228 | Nuclear pore complex protein Nup98-Nup96 ; | |
 P00973 P00973 |
OAS1_HUMAN | 400 | 12 | (2-5')oligo(A) synthase 1;2-5A synthase 1; | 2'-5'-oligoadenylate synthase 1; E18/E16;p46/p42 OAS; |
 Q9H074 Q9H074 |
PAIP1_HUMAN | 479 | 9 | PABP-interacting protein 1;PAIP-1;Poly(A)-binding protein-interacting protein 1; | Polyadenylate-binding protein-interacting protein 1 ; |
 Q8N3R9 Q8N3R9 |
PALS1_HUMAN | 675 | 230 | Protein PALS1 ; MAGUK p55 subfamily member 5;Membrane protein, palmitoylated 5 ;Protein associated with Lin-7 1 {ECO:0000303|PubMed:12527193, ECO:0000312|HGNC:HGNC:18669}; | |
 P35232 P35232 |
PHB1_HUMAN | 272 | 88 | Prohibitin 1 ; | |
 Q99623 Q99623 |
PHB2_HUMAN | 299 | 88 | Prohibitin-2; B-cell receptor-associated protein BAP37;D-prohibitin;Repressor of estrogen receptor activity; | |
 Q8NEB9 Q8NEB9 |
PK3C3_HUMAN | 887 | 616 | PI3-kinase type 3;PI3K type 3;PtdIns-3-kinase type 3; | Phosphatidylinositol 3-kinase catalytic subunit type 3; Phosphatidylinositol 3-kinase p100 subunit;Phosphoinositide-3-kinase class 3;hVps34; |
 P62937 P62937 |
PPIA_HUMAN | 165 | 761 | PPIase A; | Peptidyl-prolyl cis-trans isomerase A; Cyclophilin A;Cyclosporin A-binding protein;Rotamase A; |
 P51149 P51149 |
RAB7A_HUMAN | 207 | 1500 | Ras-related protein Rab-7a ; | |
 P05109 P05109 |
S10A8_HUMAN | 93 | 453 | CFAG;MRP-8;p8; | Protein S100-A8 ; Calgranulin-A;Calprotectin L1L subunit;Cystic fibrosis antigen;Leukocyte L1 complex light chain;Migration inhibitory factor-related protein 8;S100 calcium-binding protein A8;Urinary stone protein band A; |
 O43765 O43765 |
SGTA_HUMAN | 313 | 337 | UBP; | Small glutamine-rich tetratricopeptide repeat-containing protein alpha; Alpha-SGT;Vpu-binding protein; |
 P84022 P84022 |
SMAD3_HUMAN | 425 | 91 | MAD homolog 3;Mad3;Mothers against DPP homolog 3;hMAD-3; SMAD 3;Smad3;hSMAD3; | Mothers against decapentaplegic homolog 3; JV15-2;SMAD family member 3; |
 O95721 O95721 |
SNP29_HUMAN | 258 | 14 | SNAP-29 ; | Synaptosomal-associated protein 29 ; Soluble 29 kDa NSF attachment protein ;Vesicle-membrane fusion protein SNAP-29; |
 P56962 P56962 |
STX17_HUMAN | 302 | 9 | Syntaxin-17 ; | |
 Q9UHD2 Q9UHD2 |
TBK1_HUMAN | 729 | 1500 | Serine/threonine-protein kinase TBK1 ; NF-kappa-B-activating kinase ;T2K;TANK-binding kinase 1 ; | |
 Q8IUC6 Q8IUC6 |
TCAM1_HUMAN | 712 | 29 | TICAM-1; MyD88-3;TIR domain-containing adapter protein inducing IFN-beta; | TIR domain-containing adapter molecule 1; Proline-rich, vinculin and TIR domain-containing protein B;Putative NF-kappa-B-activating protein 502H;Toll-interleukin-1 receptor domain-containing adapter protein inducing interferon beta; |
 O15455 O15455 |
TLR3_HUMAN | 904 | 954 | Toll-like receptor 3 ; | |
 Q9NYK1 Q9NYK1 |
TLR7_HUMAN | 1049 | 832 | Toll-like receptor 7 ; | |
 Q9NR97 Q9NR97 |
TLR8_HUMAN | 1041 | 712 | Toll-like receptor 8 ; | |
 Q5BJD5 Q5BJD5 |
TM41B_HUMAN | 291 | 0 | Transmembrane protein 41B ; Protein stasimon ; | |
 O15393 O15393 |
TMPS2_HUMAN | 492 | 1500 | Transmembrane protease serine 2 ; Serine protease 10 ; | |
 Q9NRS4 Q9NRS4 |
TMPS4_HUMAN | 437 | 1500 | CAPH2;MT-SP2; | Transmembrane protease serine 4 ; Channel-activating protease 2;Membrane-type serine protease 2; |
 O94826 O94826 |
TOM70_HUMAN | 608 | 201 | Mitochondrial import receptor subunit TOM70 ; Mitochondrial precursor proteins import receptor;Translocase of outer membrane 70 kDa subunit;Translocase of outer mitochondrial membrane protein 70 ; | |
 Q8NHX9 Q8NHX9 |
TPC2_HUMAN | 752 | 176 | Two pore channel protein 2; Two pore calcium channel protein 2; | |
 P29597 P29597 |
TYK2_HUMAN | 1187 | 1500 | Non-receptor tyrosine-protein kinase TYK2; | |
 P47901 P47901 |
V1BR_HUMAN | 424 | 852 | V1bR; | Vasopressin V1b receptor ; AVPR V1b;AVPR V3;Antidiuretic hormone receptor 1b;Vasopressin V3 receptor; |
 Q9BV40 Q9BV40 |
VAMP8_HUMAN | 100 | 58 | VAMP-8 ; EDB ; | Vesicle-associated membrane protein 8 ; Endobrevin ; |
 Q96JC1 Q96JC1 |
VPS39_HUMAN | 886 | 4 | hVam6p; | Vam6/Vps39-like protein ; TRAP1-like protein; |
 P49754 P49754 |
VPS41_HUMAN | 854 | 0 | Vacuolar protein sorting-associated protein 41 homolog; S53; | |
 Q5W0Z9 Q5W0Z9 |
ZDH20_HUMAN | 365 | 12 | DHHC20 ; | Palmitoyltransferase ZDHHC20 ; Acyltransferase ZDHHC20 ;DHHC domain-containing cysteine-rich protein 20 ;Zinc finger DHHC domain-containing protein 20 ; |
 Q9ULC8 Q9ULC8 |
ZDHC8_HUMAN | 765 | 12 | DHHC-8 ; | Palmitoyltransferase ZDHHC8 ; Zinc finger DHHC domain-containing protein 8 ;Zinc finger protein 378; |
 Q9Y397 Q9Y397 |
ZDHC9_HUMAN | 364 | 12 | DHHC-9;DHHC9; | Palmitoyltransferase ZDHHC9; Zinc finger DHHC domain-containing protein 9;Zinc finger protein 379;Zinc finger protein 380; |
| protein_id | length (amino acids) |
Number of 3D homologues |
product | gene | note |
 YP_009724389.1 YP_009724389.1 |
7096 | 1500 | 2'-O-ribose methyltransferase | orf1ab | pp1ab; translated by -1 ribosomal frameshift |
 YP_009725297.1 YP_009725297.1 |
180 | 40 | leader protein | nsp1; produced by both pp1a and pp1ab | |
 YP_009725298.1 YP_009725298.1 |
638 | 8 | nsp2 | produced by both pp1a and pp1ab | |
 YP_009725299.1 YP_009725299.1 |
1945 | 1500 | nsp3 | former nsp1; conserved domains are: N-terminal | |
 YP_009725300.1 YP_009725300.1 |
500 | 24 | nsp4 | nsp4B_TM; contains transmembrane domain 2 (TM2); | |
 YP_009725301.1 YP_009725301.1 |
306 | 1500 | 3C-like proteinase | nsp5A_3CLpro and nsp5B_3CLpro; main proteinase | |
 YP_009725302.1 YP_009725302.1 |
290 | 0 | nsp6 | nsp6_TM; putative transmembrane domain; produced by | |
 YP_009725303.1 YP_009725303.1 |
83 | 117 | nsp7 | produced by both pp1a and pp1ab | |
 YP_009725304.1 YP_009725304.1 |
198 | 189 | nsp8 | produced by both pp1a and pp1ab | |
 YP_009725305.1 YP_009725305.1 |
113 | 72 | nsp9 | ssRNA-binding protein; produced by both pp1a and | |
 YP_009725306.1 YP_009725306.1 |
139 | 203 | nsp10 | nsp10_CysHis; formerly known as growth-factor-like | |
 YP_009725307.1 YP_009725307.1 |
932 | 91 | RNA-dependent RNA polymerase | nsp12; NiRAN and RdRp; produced by pp1ab only | |
 YP_009725308.1 YP_009725308.1 |
601 | 196 | helicase | nsp13_ZBD, nsp13_TB, and nsp_HEL1core; zinc-binding | |
 YP_009725309.1 YP_009725309.1 |
527 | 122 | 3'-to-5' exonuclease | nsp14A2_ExoN and nsp14B_NMT; produced by pp1ab | |
 YP_009725310.1 YP_009725310.1 |
346 | 213 | endoRNAse | nsp15-A1 and nsp15B-NendoU; produced by pp1ab only | |
 YP_009725311.1 YP_009725311.1 |
298 | 98 | 2'-O-ribose methyltransferase | nsp16_OMT; 2'-o-MT; produced by pp1ab only | |
 YP_009725295.1 YP_009725295.1 |
4405 | 1500 | nsp11 | orf1ab | pp1a |
 YP_009725312.1 YP_009725312.1 |
13 | 0 | nsp11 | produced by pp1a only | |
 YP_009724390.1 YP_009724390.1 |
1273 | 1500 | surface glycoprotein | S | structural protein; spike protein |
 YP_009724391.1 YP_009724391.1 |
275 | 12 | ORF3a protein | ORF3a | |
 YP_009724392.1 YP_009724392.1 |
75 | 18 | envelope protein | E | ORF4; structural protein; E protein |
 YP_009724393.1 YP_009724393.1 |
222 | 20 | membrane glycoprotein | M | ORF5; structural protein |
 YP_009724394.1 YP_009724394.1 |
61 | 0 | ORF6 protein | ORF6 | |
 YP_009724395.1 YP_009724395.1 |
121 | 4 | ORF7a protein | ORF7a | |
 YP_009725318.1 YP_009725318.1 |
43 | 0 | ORF7b | ORF7b | |
 YP_009724396.1 YP_009724396.1 |
121 | 9 | ORF8 protein | ORF8 | |
 YP_009724397.2 YP_009724397.2 |
419 | 365 | nucleocapsid phosphoprotein | N | ORF9; structural protein |
 YP_009725255.1 YP_009725255.1 |
38 | 0 | ORF10 protein | ORF10 |
 "Searching Contact Proteins with Query Compound" of HOMCOS for each compound. Bound protein 3D structures
"Searching Contact Proteins with Query Compound" of HOMCOS for each compound. Bound protein 3D structures  with the compound and its analogues are summarized.
with the compound and its analogues are summarized.
| KEGG_ID | structure | NAME | Number of identical compounds |
Number of similar compounds (tanimoto=0.5) |
OTHER NAME | EFFICACY |
 D01703 D01703 |
D01703 Ciclesonide (JAN/USAN/INN); Alvesco (TN); Omnaris (TN); Zetonna (TN) | 0 | 14 | Alvesco (TN); Omnaris (TN); Zetonna (TN) | Antiasthmatic, Anti-inflammatory, Glucocorticoid receptor agonist | |
 D01425 D01425 |
D01425 Lopinavir (JAN/USP/INN) | 1 | 41 | Antiviral, HIV protease inhibitor | ||
 D00427 D00427 |
D00427 Ritonavir (JAN/USP/INN); Norvir (TN) | 1 | 32 | Norvir (TN) | Antiviral, HIV protease inhibitor | |
 D09537 D09537 |
D09537 Favipiravir (JAN/USAN/INN); Avigan (TN) | 0 | 72 | Avigan (TN) | Antiviral, RNA replicase inhibitor | |
 D09537_1 D09537_1 |
D09537_1 Favipiravir-MTP; Active form of Favipiravir; | 1 | 150 | Active form of Favipiravir; | Antiviral, RNA replicase inhibitor | |
 D09537_2 D09537_2 |
D09537_2 Favipiravir-DTP; Active form of Favipiravir; | 0 | 140 | Active form of Favipiravir; | Antiviral, RNA replicase inhibitor | |
 D09537_3 D09537_3 |
D09537_3 Favipiravir-RTP; Active form of Favipiravir; | 1 | 150 | Active form of Favipiravir; | Antiviral, RNA replicase inhibitor | |
 D11472 D11472 |
D11472 Remdesivir (JAN/USAN); Veklury (TN) | 0 | 4 | Veklury (TN) | Antiviral, RNA polymerase inhibitor | |
 D11472_1 D11472_1 |
D11472_1 Remdesivir-monophosphate (USAN | 1 | 149 | Antiviral | ||
 D11472_2 D11472_2 |
D11472_2 Remdesivir-diphosphate (USAN | 0 | 138 | Antiviral | ||
 D11472_3 D11472_3 |
D11472_3 Remdesivir-triphosphate (USAN | 1 | 150 | Antiviral | ||
 D02366 D02366 |
D02366 Chloroquine (USP/INN) | 2 | 33 | Antimalarial, Amebicide | ||
 D08050 D08050 |
D08050 Hydroxychloroquine (INN); Polirreumin (TN) | 1 | 23 | Polirreumin (TN) | Antimalarial | |
 D01670 D01670 |
D01670 Nafamostat mesylate (USAN); Nafamostat mesilate (JP18); Ronastat (TN) | 1 | 35 | Nafamostat mesilate (JP18); Ronastat (TN) | Anticoagulant, Antifibrinolytic, Serine protease inhibitor | |
 D00429 D00429 |
D00429 Saquinavir (JAN/USP/INN); Fortovase (TN) | 1 | 6 | Fortovase (TN) | Antiviral, HIV protease inhibitor | |
 D01035 D01035 |
D01035 Cepharanthine (JAN); Cepharanthine (TN) | 0 | 1 | Cepharanthine (TN) | Antiallergic, Blood circulation promotor | |
 D00804 D00804 |
D00804 Ivermectin (JAN/USP/INN); Sklice (TN); Soolantra (TN); Stromectol (TN) | 0 | 2 | Sklice (TN); Soolantra (TN); Stromectol (TN) | Antiparasitic | |
 D00292 D00292 |
D00292 Dexamethasone (JP18/USP/INN); Decadron (TN); Maxidex (TN) | 1 | 116 | Decadron (TN); Maxidex (TN) | Anti-inflammatory, Antipruritic, Glucocorticoid receptor agonist | |
 D07486 D07486 |
D07486 Azithromycin (INN); Azasite (TN); Azithromycin (TN) | 1 | 26 | Azasite (TN); Azithromycin (TN) | Antibacterial, Protein biosynthesis inhibitor | |
 D08897 D08897 |
D08897 Dapagliflozin (USAN/INN); Forxiga (TN) | 1 | 39 | Forxiga (TN) | Antidiabetic, SGLT-2 inhibitor | |
 D00570 D00570 |
D00570 Colchicine (JP18/USP); Colchicine (TN) | 1 | 11 | Colchicine (TN) | Gout suppressant, Leukocyte (neutrophil) migration inhibitor, Tubulin polymerization inhibitor | |
 D07606 D07606 |
D07606 Camostat (INN) | 0 | 14 | Anti-inflammatory, Serine protease inhibitor | ||
 D10308 D10308 |
D10308 Baricitinib (JAN/USAN/INN); Olumiant (TN) | 1 | 9 | Olumiant (TN) | Anti-inflammatory, Antirheumatic, Immunosuppressant, Janus kinase (JAK) inhibitor | |
 D11943 D11943 |
D11943 Molnupiravir (JAN/USAN); Lagevrio (TN) | 1 | 115 | Lagevrio (TN) | Antiviral | |
 D12244 D12244 |
D12244 Nirmatrelvir (JAN/USAN); PF-07321332 | 3 | 81 | PF-07321332 | Antiviral | |
 D03506 D03506 |
D03506 Cinanserin hydrochloride (USAN) | 0 | 33 | Antiviral |
HOMCOS is developed and maintained by Protein Data Bank Japan, IPR, Osaka University. It is now supported by the project "Platform for Drug Discovery, Informatics, and Structural Life Science" by MEXT, Japan.
References