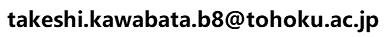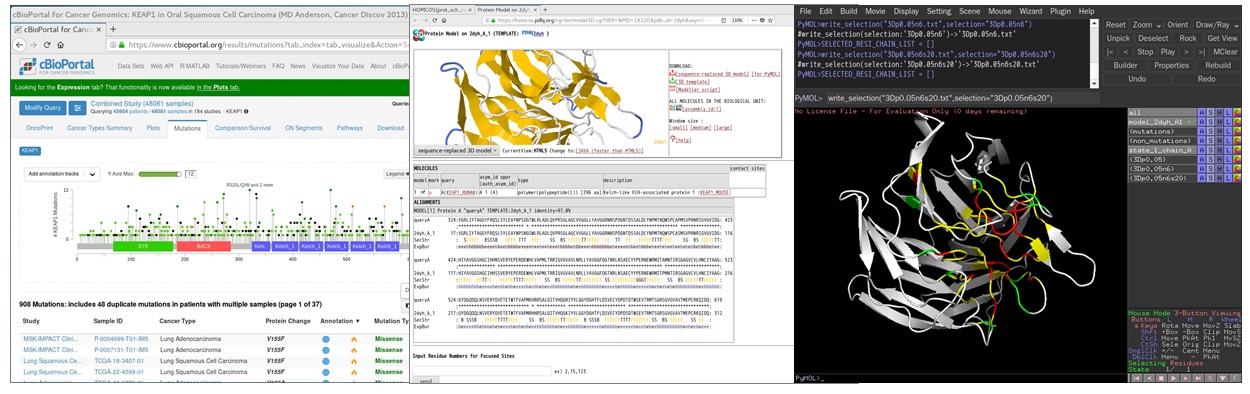
Many mutations are observed in cancer cells, it is important to extract functionally significant sites among them. This page explains, how to identify potential cancer-associated sites by "3D cluster" sites (dense mutations site in 3D space) with the help of HOMCOS server and PyMOL script py3Dcluster.py. The algorithm for detecting 3D cluster is based on Gao et al, 2017.


>PyMOL), type read_mutations([mutation_file]). In this case, [mutation_file] is "KEAP1_ProteinChange.tsv". Type read_mutations("KEAP1_ProteinChange.tsv")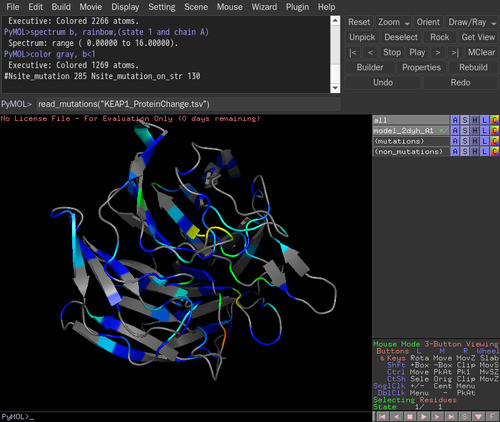
>PyMOL), type cal_3Dcluster(). Count mutations of 3D cluster is defined as the sum of count mutations of contacting residues. Two residues are regarded in contact if any pair of their atoms is within 5 angstrom.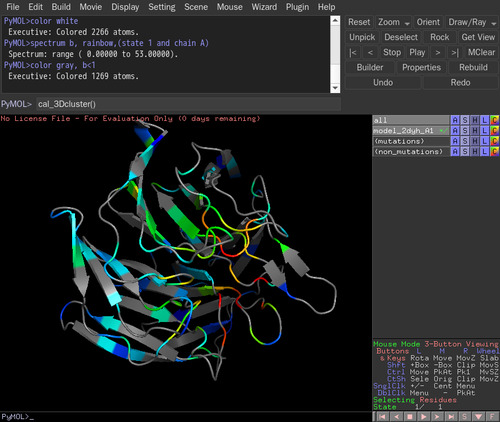
cal_3Dpvalue() in the command line. Residue numbers are randomly shuffled N times,the count mutations of 3D cluster are calculated using each shuffled residue numbers. The p-value is the fraction of the shufflings whose count mutation of 3D cluster is over the original value. The value N is 100000 in default. The value N can be changed by typing cal_3Dvalue(N=100).
cal_asa() in the command line.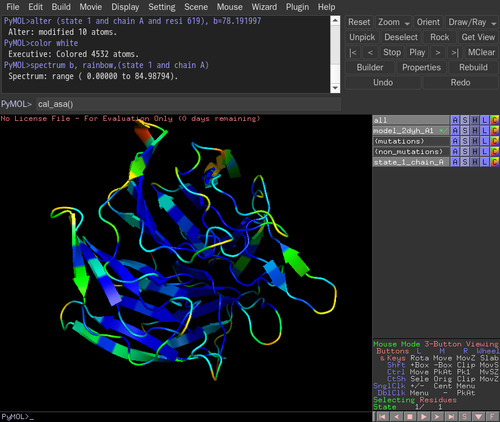
select_3Dpvalue() in the command line. The default p-value is 0.05. The low p-value residues are named as "3Dp0.05". The p-value can be changed, such as select_3Dpvalue(p=0.01). 
select_neighbor(selection="3Dp0.05") in the command line. The default distance threshold is 6 angstrom. The selected residues are named as "3Dp0.05n6". The distance threshold can be changed, such as select_neighbor(selection="3Dp0.05",Dthre=8). 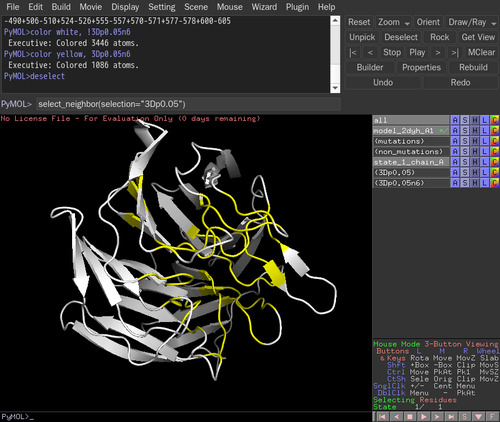
select_neighbor_surf(selection="3Dp0.05") in the command line. The default relative ASA(%) is 20 %. The selected residues are named as "3Dp0.05n6s20". The relative ASA(%) can be changed, such as select_neighbor_surf(selection="3Dp0.05",relasa=30). 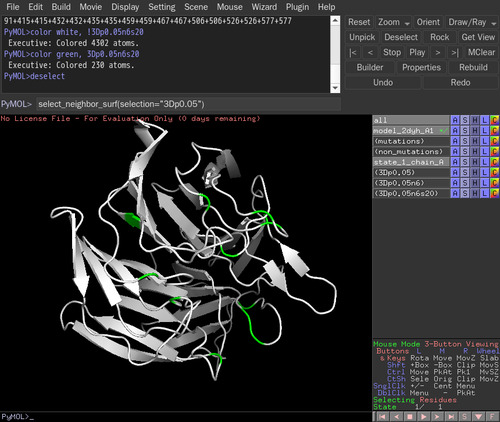
write_selection("3Dp0.05n6s20.txt",selection="3Dp0.05n6s20") in the command line. The output file is "3Dp0.05n6s20.txt". Similarly, other selected residues can be saved such as write_selection("3Dp0.05.txt",selection="3Dp0.05") and write_selection("3Dp0.05n6.txt",selection="3Dp0.05n6"). 
>> 3Dp0.05n6s20.txt <<
#write_selection(ofname='3Dp0.05n6s20.txt',selection='3Dp0.05n6s20') #COMMAND pymol_3Dcluster.py #DATE 2021/05/16 14:13:55 #MIN_RELASA 20.000000 MIN_MINDIS_FRM_SELECTED 6.000000 #FOCUS_MODEL 1 FOCUS_CHAIN A #[chain:1] [resi:2] [resn:3] #[MINDIS_FRM_SELECTED(A):4] [RELASA(%):5] #[count_mutation:6] [cluster3D(mutation_density):7] #[pvalue_cluster3D:8] [-log(pvalue_cluster3D):9] #[contact_residues:10] A 334 TYR 20.33 3.78 0 0 -1.000000 -1.000000 332 333 335 336 337 338 339 363 382 577 600 601 602 603 A 335 PHE 29.51 5.20 1 26 +0.081920 +1.086610 333 334 336 337 339 577 600 601 A 341 TYR 20.68 5.15 0 0 -1.000000 -1.000000 330 331 332 339 340 342 343 354 357 358 599 601 A 391 SER 22.19 5.95 0 0 -1.000000 -1.000000 378 389 390 392 393 408 409 411 412 413 A 415 ARG 21.00 0.00 6 50 +0.002120 +2.673664 364 379 413 414 416 417 430 431 461 462 463 508 509 556 A 432 HIS 32.80 3.81 0 0 -1.000000 -1.000000 389 411 412 430 431 433 434 435 437 A 435 ILE 42.32 5.62 0 0 -1.000000 -1.000000 430 431 432 433 434 436 437 461 A 459 ARG 47.35 3.82 1 17 +0.297920 +0.525900 436 437 438 456 457 458 460 461 478 479 484 A 467 VAL 23.18 4.59 0 0 -1.000000 -1.000000 419 420 421 465 466 468 469 470 471 472 473 514 A 506 ILE 20.19 3.81 1 13 +0.472110 +0.325957 483 484 485 503 504 505 507 508 525 526 531 A 526 ASP 22.43 4.97 0 0 -1.000000 -1.000000 483 505 506 507 524 525 527 528 529 531 A 577 PHE 20.52 5.05 0 0 -1.000000 -1.000000 334 335 571 572 575 576 578 579 600 601 602 A 334 TYR 20.33 3.78 0 0 -1.000000 -1.000000 332 333 335 336 337 338 339 363 382 577 600 601 602 603 A 335 PHE 29.51 5.20 1 26 +0.081920 +1.086610 333 334 336 337 339 577 600 601 A 341 TYR 20.68 5.15 0 0 -1.000000 -1.000000 330 331 332 339 340 342 343 354 357 358 599 601 A 391 SER 22.19 5.95 0 0 -1.000000 -1.000000 378 389 390 392 393 408 409 411 412 413 A 415 ARG 21.00 0.00 6 50 +0.002120 +2.673664 364 379 413 414 416 417 430 431 461 462 463 508 509 556 A 432 HIS 32.80 3.81 0 0 -1.000000 -1.000000 389 411 412 430 431 433 434 435 437 A 435 ILE 42.32 5.62 0 0 -1.000000 -1.000000 430 431 432 433 434 436 437 461 A 459 ARG 47.35 3.82 1 17 +0.297920 +0.525900 436 437 438 456 457 458 460 461 478 479 484 A 467 VAL 23.18 4.59 0 0 -1.000000 -1.000000 419 420 421 465 466 468 469 470 471 472 473 514 A 506 ILE 20.19 3.81 1 13 +0.472110 +0.325957 483 484 485 503 504 505 507 508 525 526 531 A 526 ASP 22.43 4.97 0 0 -1.000000 -1.000000 483 505 506 507 524 525 527 528 529 531 A 577 PHE 20.52 5.05 0 0 -1.000000 -1.000000 334 335 571 572 575 576 578 579 600 601 602
mk_dic_resn_frm_resi_from_get_model()
translate_mutation_resi_list_from_string(sites_string)
make_resi_list_on_structure(resi_list)
make_pymol_select_str_from_resi_list(resi_list)
mk_decimal_string(x)
random_permutation_list(array)
sort_num_str_list(str_list)
LastModfied:2023/01/31
Questions and comments: