|
|
| PID | QueryLength | FocusSite | TITLE |
| 9058 | 422 | 271 V | RecName: Full=Nucleoprotein ;AltName: Full=Nucleocapsid protein ; Short=NC ; Short=Protein N ; |
| AC/ID | AC:P59595 ID:NCAP_SARS |
| Feature Table for 271-th site |
HELIX: COMPBIAS: /note="Polar residues" REGION: /note="Disordered" REGION: /note="Dimerization" DOMAIN: /note="CoV N CTD" CHAIN: /note="Nucleoprotein" /id="PRO_0000106003" |
Percentages of Amino Acids in Homologous Proteins
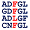 [show alignment] [show alignment]
|
V:88% L:6% I:4% M:3% |
| Template For Monomer | predicted SecStr | predicted ExpBur | Predicted Relative Acc(%) |
 8fg2 8fg2 |
H (alpha-helix) | b (buried) | 3.3 |
| Predicted Bind Molecules |
| homo:19 precipitant:2 |
| Templates for 3D complexes |
homo [26763:Q4ZJS4_9CORO
]  2ca1_A_1_B_1 2ca1_A_1_B_1  2ge7_A_1_B_1 2ge7_A_1_B_1  2ge7_B_1_A_1 2ge7_B_1_A_1  2ge8_A_1_B_1 2ge8_A_1_B_1  2ge8_B_1_A_1 2ge8_B_1_A_1  2ge8_C_1_D_1 2ge8_C_1_D_1  2ge8_D_1_C_1 2ge8_D_1_C_1  2ge8_E_1_F_1 2ge8_E_1_F_1  2ge8_F_1_E_1 2ge8_F_1_E_1  2ge8_G_1_H_1 2ge8_G_1_H_1  2ge8_H_1_G_1 [33389:NCAP_CVHNL
] 2ge8_H_1_G_1 [33389:NCAP_CVHNL
]  5epw_A_1_B_1 5epw_A_1_B_1  5epw_B_1_A_1 [32372:A0A0D3MU51_9BETC
] 5epw_B_1_A_1 [32372:A0A0D3MU51_9BETC
]  6g13_A_1_C_1 6g13_A_1_C_1  6g13_B_1_D_1 6g13_B_1_D_1  6g13_C_1_A_1 6g13_C_1_A_1  6g13_D_1_B_1 [23845:NCAP_SARS2
] 6g13_D_1_B_1 [23845:NCAP_SARS2
]  7xwx_D_1_C_1 7xwx_D_1_C_1  7xwx_F_1_B_1
precipitant [ACT ] 7xwx_F_1_B_1
precipitant [ACT ]  7n0i_D_1_M_1 [GOL ] 7n0i_D_1_M_1 [GOL ]  7o36_B_1_F_1 7o36_B_1_F_1
|
 [Back to SiteTable] |
 [Back to Search Page] |
 [Back to Top Page] |