|
|
| PID | QueryLength | FocusSite | TITLE |
| 676673 | 805 | 135 P | RecName: Full=Angiotensin-converting enzyme 2; EC=3.4.17.23 ;AltName: Full=Angiotensin-converting enzyme homolog ; Short=ACEH ;AltName: Full=Angiotensin-converting enzyme-related carboxypeptidase ; Short=ACE-related carboxypeptidase; EC=3.4.17.- ;AltName: Full=Metalloprotease MPROT15 ;Contains: RecName: Full=Processed angiotensin-converting enzyme 2 ;Flags: Precursor; |
| AC/ID | AC:Q9BYF1 ID:ACE2_HUMAN |
| Feature Table for 135-th site |
MUTAGEN: /note="PD->SM: No effect on interaction with SARS-CoV spike glycoprotein." VAR_SEQ: /note="MSSSSWLLLSLVAVTAAQSTIEEQAKTFLDKFNHEAEDLFYQSSLASWNYNT NITEENVQNMNNAGDKWSAFLKEQSTLAQMYPLQEIQNLTVKLQLQALQQNGSSVLSED KSKRLNTILNTMSTIYSTGKVCNPDNPQECLLLEPGLNEIMANSLDYNERLWAWESWRS EVGKQLRPLYEEYVVLKNEMARANHYEDYGDYWRGDYEVNGVDGYDYSRGQLIEDVEHT FEEIKPLYEHLHAYVRAKLMNAYPSYISPIGCLPAHLLGDMWGRFWTNLYSLTVPFGQK PNIDVTDAMVDQAWDAQRIFKEAEKFFVSVGLPNMTQGFWENSMLTDPGNVQKAVCHPT AWDLGKGDF -> MREAGWDKGG (in isoform 2)" /id="VSP_060903" DOMAIN: /note="Peptidase M2" CHAIN: /note="Processed angiotensin-converting enzyme 2" /id="PRO_0000292268" TOPO_DOM: /note="Extracellular" CHAIN: /note="Angiotensin-converting enzyme 2" /id="PRO_0000028570" |
Percentages of Amino Acids in Homologous Proteins
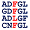 [show alignment] [show alignment]
|
P:35% Y:20% K:14% L:8% D:7% T:6% A:4% S:3% V:3% |
| Template For Monomer | predicted SecStr | predicted ExpBur | Predicted Relative Acc(%) |
 7v61 7v61 |
S (bend) | e (exposed) | 35.7 |
| Predicted Bind Molecules |
| hetero:1 |
| Templates for 3D complexes |
hetero [24941 ]  1r4l_A_1_C_1 1r4l_A_1_C_1
|
 [Back to SiteTable] |
 [Back to Search Page] |
 [Back to Top Page] |