|
|
| PID | QueryLength | FocusSite | TITLE |
| 2692367 | 631 | 602 R | RecName: Full=Sphingomyelin phosphodiesterase ; EC=3.1.4.12 ; EC=3.1.4.3 ;AltName: Full=Acid sphingomyelinase ; Short=aSMase;Contains: RecName: Full=Sphingomyelin phosphodiesterase, processed form ;Flags: Precursor; |
| AC/ID | AC:P17405 ID:ASM_HUMAN |
| Feature Table for 602-th site |
VARIANT: /note="R -> H (in NPDB and NPDA; expresses protein level comparable to wild-type SMPD1 expressing cells; retains about 10% residual enzyme activity; loss of location to lysosome; dbSNP:rs370129081)" ECO:0000269|PubMed:16010684, ECO:0000269|PubMed:21098024, ECO:0000269|PubMed:27338287" /id="VAR_060932" VARIANT: /note="R -> P (in NPDB; expresses protein level comparable to wild-type SMPD1 expressing cells; retains very low enzyme activity; loss of location to lysosome)" ECO:0000269|PubMed:16010684, ECO:0000269|PubMed:21098024" /id="VAR_060933" STRAND: CHAIN: /note="Sphingomyelin phosphodiesterase, processed form" /id="PRO_0000456684" CHAIN: /note="Sphingomyelin phosphodiesterase" /id="PRO_0000002323" |
| VARIANT for 602-th site |
R->H LP/P dbSNP:rs3701290 " Niemann-Pick disease A (NPDA) " [MIM:257200] R->H LP/P dbSNP:rs3701290 " Niemann-Pick disease B (NPDB) " [MIM:607616] R->P LP/P " Niemann-Pick disease B (NPDB) " [MIM:607616] |
Percentages of Amino Acids in Homologous Proteins
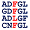 [show alignment] [show alignment]
|
R:31% D:29% A:20% H:13% M:7% |
| Template For Monomer | predicted SecStr | predicted ExpBur | Predicted Relative Acc(%) |
 5jg8 5jg8 |
S (bend) | b (buried) | 7.9 |
| Predicted Bind Molecules |
| precipitant:2 |
| Templates for 3D complexes |
precipitant [SO4 ]  5fib_A_1_R_1 [PO4 ] 5fib_A_1_R_1 [PO4 ]  5fic_A_1_O_1 5fic_A_1_O_1
|
 [Back to SiteTable] |
 [Back to Search Page] |
 [Back to Top Page] |