|
|
| PID | QueryLength | FocusSite | TITLE |
| 14620 | 1255 | 502 E | RecName: Full=Spike glycoprotein ; Short=S glycoprotein ;AltName: Full=E2 ;AltName: Full=Peplomer protein ;Contains: RecName: Full=Spike protein S1 ;Contains: RecName: Full=Spike protein S2 ;Contains: RecName: Full=Spike protein S2' ;Flags: Precursor; |
| AC/ID | AC:P59594 ID:SPIKE_SARS |
| Feature Table for 502-th site |
DOMAIN: /note="BetaCoV S1-CTD" REGION: /note="Receptor-binding domain (RBD)" CHAIN: /note="Spike protein S1" /id="PRO_0000037209" TOPO_DOM: /note="Extracellular" CHAIN: /note="Spike glycoprotein" /id="PRO_0000037208" |
Percentages of Amino Acids in Homologous Proteins
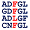 [show alignment] [show alignment]
|
I:24% E:21% S:16% K:12% F:10% G:9% Q:8% |
| Template For Monomer | predicted SecStr | predicted ExpBur | Predicted Relative Acc(%) |
 7sg4 7sg4 |
E (beta-strand) | e (exposed) | 33.7 |
| Predicted Bind Molecules |
| homo:45 |
| Templates for 3D complexes |
homo [100768 ]  5x58_A_1_B_1 5x58_A_1_B_1  5x58_B_1_C_1 5x58_B_1_C_1  5x58_C_1_A_1 5x58_C_1_A_1  5x5b_B_1_C_1 5x5b_B_1_C_1  5x5b_C_1_A_1 5x5b_C_1_A_1  5xlr_A_1_C_1 5xlr_A_1_C_1  5xlr_B_1_A_1 5xlr_B_1_A_1  6acc_A_1_C_1 6acc_A_1_C_1  6acc_B_1_A_1 6acc_B_1_A_1  6acc_C_1_B_1 6acc_C_1_B_1  6acd_B_1_A_1 6acd_B_1_A_1  6acg_A_1_C_1 6acg_A_1_C_1  6crw_A_1_B_1 6crw_A_1_B_1  6crw_C_1_A_1 6crw_C_1_A_1  6crx_B_1_C_1 6crx_B_1_C_1  6crz_A_1_B_1 6crz_A_1_B_1  6crz_C_1_A_1 6crz_C_1_A_1  6cs0_A_1_B_1 6cs0_A_1_B_1  6cs0_C_1_A_1 6cs0_C_1_A_1  6cs1_B_1_C_1 6cs1_B_1_C_1  7akj_A_1_G_1 7akj_A_1_G_1  7akj_B_1_A_1 7akj_B_1_A_1  7akj_G_1_B_1 7akj_G_1_B_1  8h0x_A_1_B_1 8h0x_A_1_B_1  8h0x_B_1_C_1 8h0x_B_1_C_1  8h0x_C_1_A_1 8h0x_C_1_A_1  8h0y_A_1_C_1 8h0y_A_1_C_1  8h0y_B_1_A_1 8h0y_B_1_A_1  8h0y_C_1_B_1 8h0y_C_1_B_1  8h0z_C_1_B_1 8h0z_C_1_B_1  8h11_A_1_C_1 8h11_A_1_C_1  8h11_B_1_A_1 8h11_B_1_A_1  8h11_C_1_B_1 8h11_C_1_B_1  8h12_A_1_B_1 8h12_A_1_B_1  8h12_B_1_C_1 8h12_B_1_C_1  8h12_C_1_A_1 8h12_C_1_A_1  8h15_A_1_B_1 8h15_A_1_B_1  8h15_C_1_A_1 8h15_C_1_A_1  8h16_A_1_C_1 8h16_A_1_C_1  8tc1_A_1_C_1 8tc1_A_1_C_1  8tc1_B_1_A_1 8tc1_B_1_A_1  8tc1_C_1_B_1 [101243:SPIKE_SARS
:WAC_BPT4
] 8tc1_C_1_B_1 [101243:SPIKE_SARS
:WAC_BPT4
]  7zh5_A_1_C_1 [101246:SPIKE_SARS
:WAC_BPT4
] 7zh5_A_1_C_1 [101246:SPIKE_SARS
:WAC_BPT4
]  7zh5_B_1_A_1 [101265:SPIKE_SARS
] 7zh5_B_1_A_1 [101265:SPIKE_SARS
]  8h16_C_1_B_1 8h16_C_1_B_1
|
 [Back to SiteTable] |
 [Back to Search Page] |
 [Back to Top Page] |